RFP LAB:
Purpose: The purpose of this lab was to express the Red Fluorescent Protein (RFP), originally found in jellyfish, in E. Coli bacteria. In doing so, we learned all the steps of genetic engineering and were given the lab opportunity to complete the experiment hands-on.
Materials & Procedure:
Lab 2A - materials and procedure can be found in Amgen lab manual 2a
Lab 4A - materials and procedure can be found in Amgen lab manual 4a
Lab 6A - materials and procedure can be found in Amgen lab manual 6a
Lab 2A - materials and procedure can be found in Amgen lab manual 2a
Lab 4A - materials and procedure can be found in Amgen lab manual 4a
Lab 6A - materials and procedure can be found in Amgen lab manual 6a
Lab 4A: Verification of Plasmid Digest by Electrophoresis
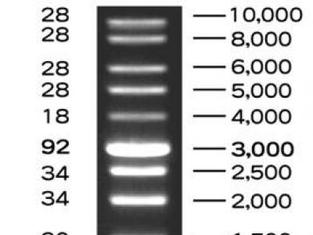
To ensure that we had actually cut the plasmid correctly, we used the process of electrophoresis. This means that we ran the plasmid in a gel alongside a DNA ladder. A DNA ladder has different-sized molecules already programmed into it, so after being run, it provides a "ladder" of bands that can be corresponded with the other lanes in order to decipher their molecule sizes. Because we already knew the size of the RFP gene and the plasmid, we were able to say if the digest had gone smoothly.
To ensure that we had actually cut the plasmid correctly, we used the process of electrophoresis. This means that we ran the plasmid in a gel alongside a DNA ladder. A DNA ladder has different-sized molecules already programmed into it, so after being run, it provides a "ladder" of bands that can be corresponded with the other lanes in order to decipher their molecule sizes. Because we already knew the size of the RFP gene and the plasmid, we were able to say if the digest had gone smoothly.
Experimental Overview:
Lab 2A: Verification of Plasmid by Restriction Digest
In this part of the lab, we cut a plasmid (a circular piece of DNA) with BamH1 and HindIII. In doing so, we were able to extract the segment of the RFP-Ara gene.
Lab 4A: Verification of Plasmid Digest by Electrophoresis
To ensure that we had actually cut the plasmid correctly, we used the process of electrophoresis. This means that we ran the plasmid in a gel alongside a DNA ladder. A DNA ladder has different-sized molecules already programmed into it, so after being run, it provides a "ladder" of bands that can be corresponded with the other lanes in order to decipher their molecule sizes. Because we already knew the size of the RFP gene and the plasmid, we were able to say if the digest had gone smoothly.
Lab 5A: Transformation of Bacteria with Recombinant Plasmid
The heart of the RFP lab was in this section, where we transformed the bacteria into a recombinant (man-made) plasmid. We used restriction enzymes to cut the plasmid and ligase to paste the gene of interest. We then used a selective marker (Amp-R, resistance to Ampicillin).
Lab 6A: Separating the RFP Gene with Column Chromatography
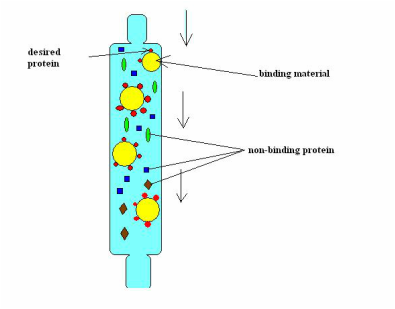
After incubating the RFP gene, we used a chromatography column in order to separate the RFP from everything else. We did this by using a column with hydrophilic beads: this effectively collected the RFP in the resin bed. Then we eluted the RFP and knocked it off those beads by using a buffer.
Lab 2A: Verification of Plasmid by Restriction Digest
In this part of the lab, we cut a plasmid (a circular piece of DNA) with BamH1 and HindIII. In doing so, we were able to extract the segment of the RFP-Ara gene.
Lab 4A: Verification of Plasmid Digest by Electrophoresis
To ensure that we had actually cut the plasmid correctly, we used the process of electrophoresis. This means that we ran the plasmid in a gel alongside a DNA ladder. A DNA ladder has different-sized molecules already programmed into it, so after being run, it provides a "ladder" of bands that can be corresponded with the other lanes in order to decipher their molecule sizes. Because we already knew the size of the RFP gene and the plasmid, we were able to say if the digest had gone smoothly.
Lab 5A: Transformation of Bacteria with Recombinant Plasmid
The heart of the RFP lab was in this section, where we transformed the bacteria into a recombinant (man-made) plasmid. We used restriction enzymes to cut the plasmid and ligase to paste the gene of interest. We then used a selective marker (Amp-R, resistance to Ampicillin).
Lab 6A: Separating the RFP Gene with Column Chromatography
After incubating the RFP gene, we used a chromatography column in order to separate the RFP from everything else. We did this by using a column with hydrophilic beads: this effectively collected the RFP in the resin bed. Then we eluted the RFP and knocked it off those beads by using a buffer.
Results:
Lab 2A & 4A: Verification of Plasmid by Restriction Digest and Electrophoresis
- We cut a plasmid, extracting the RFP-Ara gene, and ran it through a gel. There, we saw that the size of the molecule was accurate -- it corresponded with the DNA ladder.
Lab 2A & 4A: Verification of Plasmid by Restriction Digest and Electrophoresis
- We cut a plasmid, extracting the RFP-Ara gene, and ran it through a gel. There, we saw that the size of the molecule was accurate -- it corresponded with the DNA ladder.
Lab 5A: Transformation of Bacteria with Recombinant Plasmid
- We incubated three plates of bacteria in Luria broth:
- We incubated three plates of bacteria in Luria broth:
|
Plate 1:
We applied the Amp-R plasmid and ampicillin - this effectively killed off all of the plasmids that had not correctly acquired the desired gene of interest. This plate then had a culture of bacteria with the plasmid. |
Plate 2:
We applied the Amp-R plasmid but no ampicillin - because of this, the bacteria grew abundantly and without any form of selection. None died off at all, unlike Plate 1. |
Plate 3:
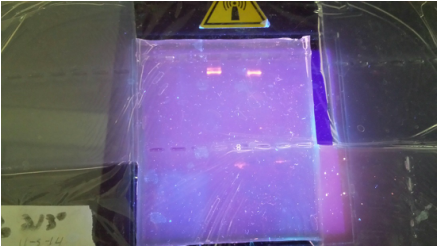
We applied the Amp-R plasmid, ampicillin, and arabinose - this killed off any bacteria that did not uptake the gene, and also expressed the red fluorescent protein. |
Lab 6A: Separating the RFP Gene with Gene Chromatography